I agreed to make my 23andMe genotyping results available publicly as part of GNZ because I knew that the results were slightly dull, and I’m not majorly at high or low risk for any diseases.
I was also very unsurprised to find out that I have blue eyes and that I was identified to be most likely descended from Northern European Ancestry.
A few hours after releasing the data, I was directed to Dienekes Pontikos’ post, where they wrote about the results of running all our data through his ancestry prediction program.
Most people were predicted to be of Northwestern European descent, and I was given an estimate of 20% Ashkenazi Jewish ancestry. When asked about this, I had no answers to give and instead decided to do my own research.

What Does Dienekes’ Program Do?
This program is based on a paper that explores the differences in allele frequencies in European Americans.
In the study, the authors identified two main components of variation in ancestry that corresponded to three groups: Mostly northwest European descent, southeast European descent, and Ashkenazi Jewish descent.
They created a list of 300 genetic markers that gave information about the ancestry in the sample, which was made publicly available.
The program uses those allele frequencies at the level of markers in the 23andMe platform to infer the proportional membership of an individual from each group.
If someone has the genotype CC at an SNP, and the C allele has 20% frequency in northwestern Europe and 60% frequency in Ashkenazi, the person is more likely to be Ashkenazi. This method put me at 20% Ashkenazi ancestry, but with not enough data to make this a confident statement.
The question left to be answered is ‘what does this mean?’ It doesn’t mean that I have 20% Ashkenazi ancestry; it means that I carry the alleles that are rare in Europe and more common in the Ashkenazi Jewish population.
However, this is based on the assumption that my ancestry only goes back to these three locations.
Knowing that I had a grandparent of Italian descent, I tested for southern European ancestry. It is also commonly known that southern European populations are similar to the Ashkenazi populations in relation to genetics.
Visualizing Population Structure with Genomes Unzipped Data
In order to explore how the GNZ participants relate to European populations, I was able to combine multiple data sources. These were from the Human Genome Diversity Panel, the 12 GNZ individuals, and a set of Ashkenazi Jewish individuals.
They were all genotyped on Illumina arrays. To begin to look at the relationships between these individuals, I used the smartpca program to look at the average genetic relationships between the populations.
In the below plot, each point is an individual on the most important axis of genetic variation in this sample. The Ashkenazi population is blue, and the GNZ individuals are black.
Dan is with the Ashkenazi population, Vincent with the French, and me with the French on component 1 and the Italians on component 2.
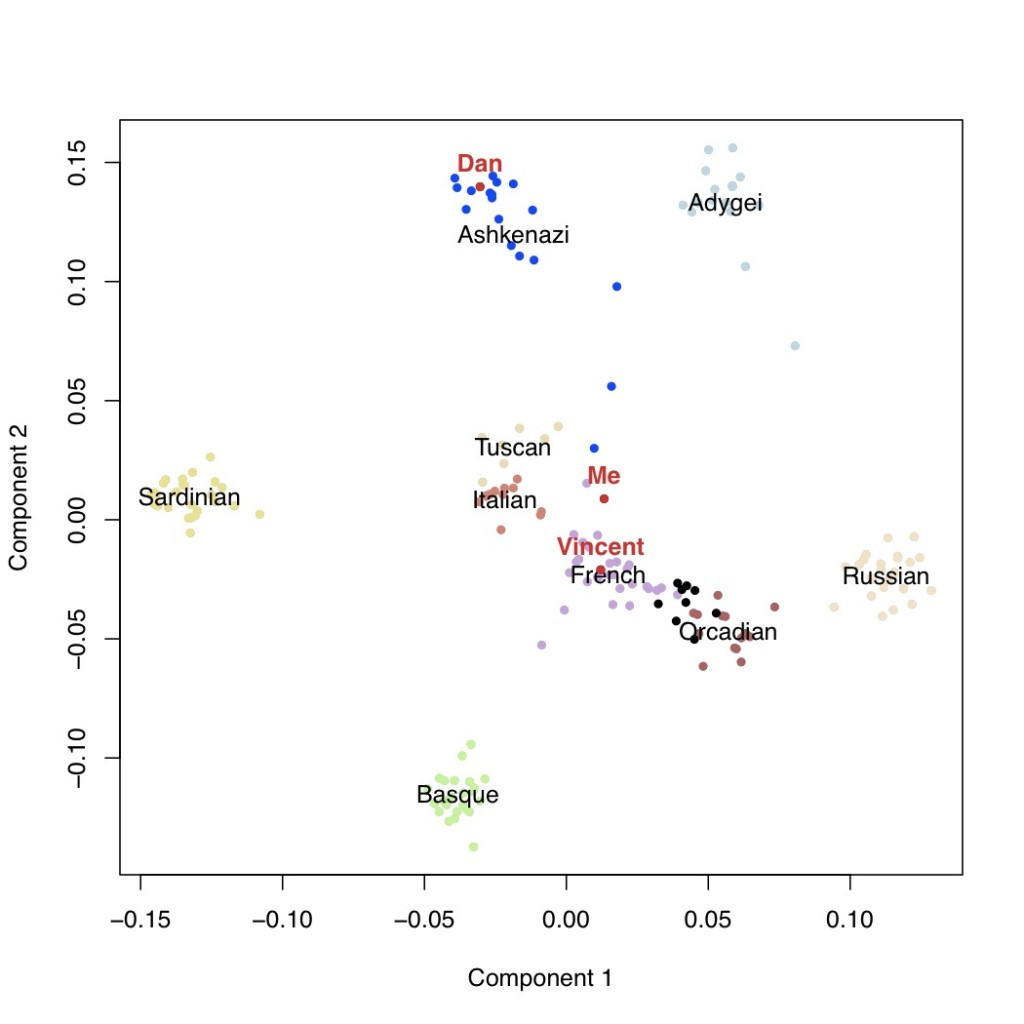
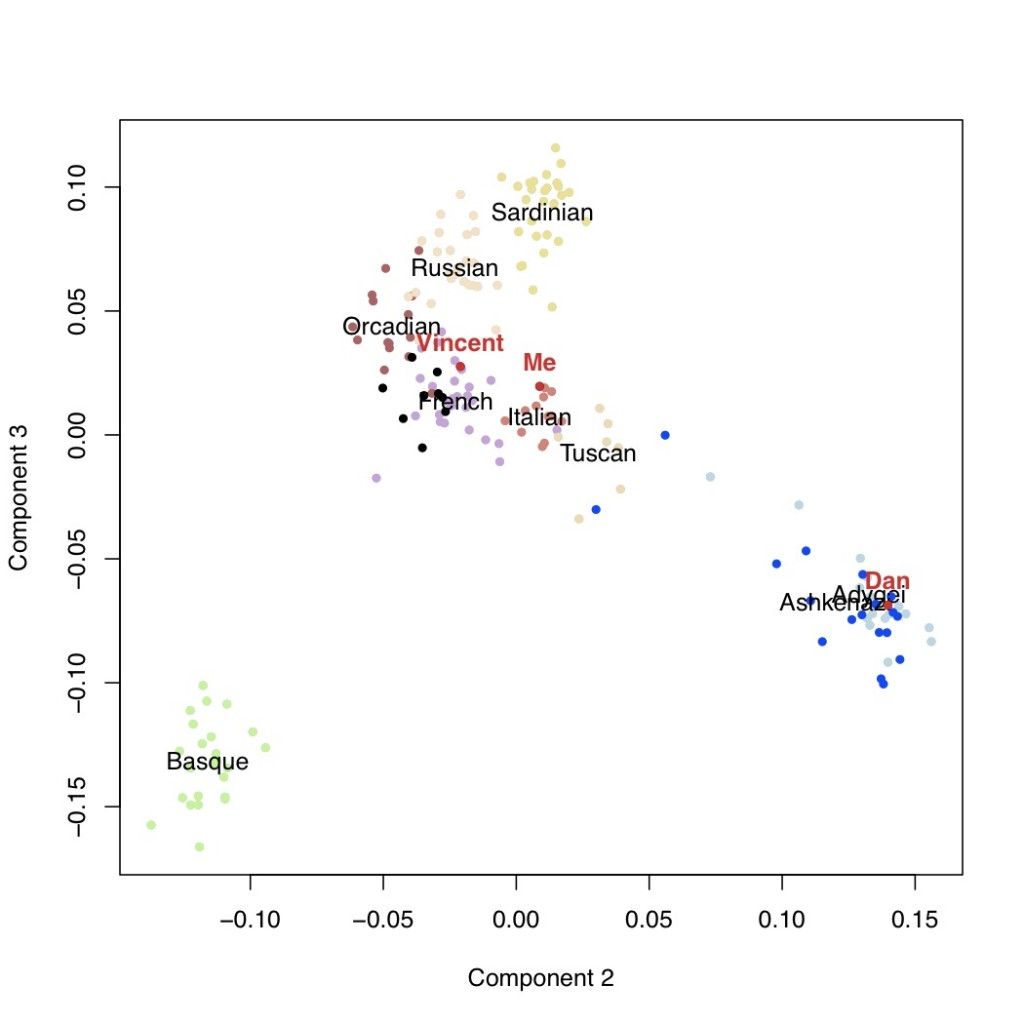
Next are the additional components of variation with the second versus third components, followed by the third versus the fourth.
The fourth axis of variation separates the Ashkenazi population from the rest of Europe.
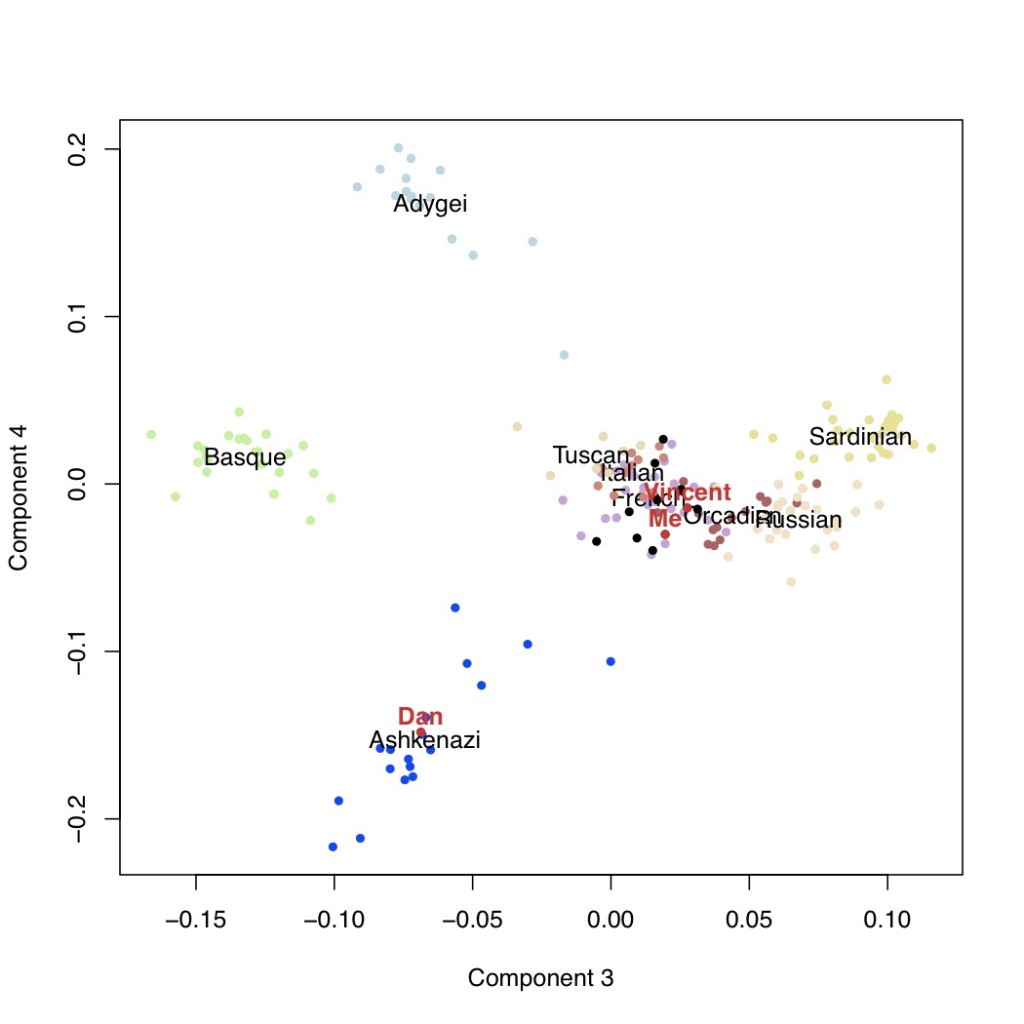
Conclusion
The analysis suggests that I do not have any Ashkenazi Jewish ancestry, and the results were actually hidden south European ancestry.
I have satisfied my curiosity surrounding my knowledge of family history, and it seems I have a small amount of south European ancestry mixed with a large amount of northwest European ancestry.
