The genome scans currently offered by major personal genomics companies provide information about only one kind of genetic variation: single nucleotide polymorphisms, or SNPs. However, SNPs are just one end of a size spectrum of variation, reaching all the way up to large duplications or deletions of DNA known as copy number variants (CNVs). Over the last decade we have learned that CNVs are a surprisingly common form of variation in humans, and they span a formidable chunk of the genome. While there are about 3M-3.5M bases of variation due to SNPs within an individual genome (in say, a typical person of European descent), there are at least 50-60M variable bases due to CNVs.
For the personal genome enthusiast with their SNP chip data from 23andMe or deCODEme in hand, there are two important practical questions: (1) can I learn about my CNVs using SNP chip data; and (2) will that information be useful?
In this post I will discuss ways to squeeze information about CNVs out of current SNP chip datasets. However, I will also argue that for the purpose of cataloging one’s own genetic variants, and especially for the purpose of understanding the complete functional consequences of one’s own genome sequence, it may well be worth waiting for whole genome sequencing.
What is a CNV?
CNVs are gains or losses of contiguous DNA sequence that can be identified by comparing multiple genomes. Strictly speaking, CNVs can range in size from 1bp to over 1Mb in size, although for historical reasons some people qualitatively divide this size spectrum into small “indels” (< 50 or 100bp) and large CNVs (everything else). The vast majority of CNVs are small.
Detecting CNVs directly from SNP chip intensity data
CNVs can be directly identified using SNP chip data by quantifying the amount of DNA hybridizing to each SNP probe: people with more copies of a particular region will have more DNA binding to that location on the chip than people with fewer copies. This can be measured by looking at the intensity of the signal at that location on the chip.
However, most CNVs are small, and the density of probes on SNP arrays is low enough that the majority of CNVs will not actually contain a probe. More importantly, personal genomics companies don’t currently provide customers with the raw data required to estimate intensity; unless this changes, customers won’t be able to use this approach on their own genome scan data.
Still, let’s say we were able to access intensity data for our genome scans – what could we find?
Our current best estimate is that there are 800,000 CNVs >= 1bp in a single genome. This number scales down to approximately 2700 when considering events >1 kb, which is the lower end of the size spectrum possible to detect with the SNP arrays used by DTC companies. The number of CNVs that are actually detected from SNP chip data will depend on the algorithm used and the quality of the experiment.
With a typical experiment and conservative analysis I would expect on average of 70-90 CNVs to be detected on Illumina 1M, and 20-40 CNVs with Illumina HumanHap-550, two of the platforms used for personal genetics (by deCODEme and 23andMe, respectively). The probes for both of these platforms were developed prior to the generation of many of the new high-resolution CNV maps, and newer Illumina SNP chips should have much better coverage of large, common CNVs.
Using your family
There is another, indirect way that CNVs can be detected from SNP chip data without any access to the raw intensity files: by tracing the patterns of inheritance of particular SNPs within your family, and looking for places where that pattern is inconsistent with normal expectations. Departures from “Mendelian” inheritance can provide clues about a CNV lurking in that region of your genome.
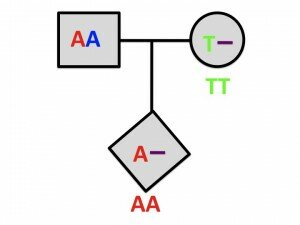
The concept behind the method is simple: SNP genotyping algorithms that are naïve to CNVs often mis-call a person who is heterozygous for a deletion as homozygous for the nucleotide that is present. What this means is that when a deletion is transmitted from parent to child, the SNP genotypes that are called at that position can give the impression that the deletion-bearing parent hasn’t transmitted any genetic material at all! This would be the case when the child inherits a base from the undeleted parent that is not present in the deleted parent. Of course, this could happen due to plain old genotyping error, so such incompatibilities need to be unusually clustered on a chromosome in order for us to be statistically confident that there is a deletion present.
Curiously, the power of this method depends on SNP density, so that families from the populations with greatest diversity will have the most success chance at finding a deletion this way. This type of analysis was first done genome-wide in 2005, when two groups used the 1 million SNPs from HapMap I to identify about 11 deletions/trio in a population of European ancestry and 20 deletions/trio in a population from Nigeria. The false discovery rate was empirically estimated to be 14% (these numbers are from the Conrad, et al. version). Based on these results I would expect the numbers to scale to around 5-10 deletions discovered per trio using a 550K SNP chip.
Indirect detection of CNVs via imputation
Many common CNVs can be assayed indirectly – or “imputed” – using your SNP genotypes. Publicly available resources make it possible to define a set of nearby SNPs that are strongly associated with a particular CNV. Then, using freely available software, one can impute (make a statistical best-guess estimate) of your CNV genotypes based on your own SNP data.
This is a statistical exercise, so a probability will be assigned to each genotype, but in many cases SNP data are informative enough to impute CNVs with high accuracy. In a recent study of Craig Venter’s genome, the authors concluded that as much as 75% of the SVs detected in his genome could have been imputed from public datasets. This is the first analysis of this nature, so the numbers may fluctuate, but I suspect that we will be able to impute common CNVs with broadly the same accuracy as common SNPs.
In the short term imputation is probably the best way to assay common CNVs in a single genome, and it is the preferred approach currently taken by most researchers performing genome-wide association studies of common traits and diseases using SNP chips: allow the direct CNV genotyping to be done by specialists in a shared resource like the HapMap samples, and then impute.
Importantly, this approach currently only works for CNVs that are present at a reasonable frequency in the population (i.e. >5%), and will not allow one to access rare CNVs or – even more interestingly – events that have occurred uniquely in your own genome rather than being inherited from your parents, so-called “de novo” CNVs. In individuals of European descent, we estimate that there are ~5,000 CNVs >1kb that are common enough to be potentially imputed with today’s resources. The soon-to-be-published 1000 genomes pilot project has generated genotypes on over 500,000 smaller CNVs (i.e. indels), many of which will now be imputable.
Do CNVs matter?
Contrary to the intuition that large polymorphisms should have large effects on traits, the impact of common CNVs on common traits studied thus far appears to be surprisingly small. Fewer than 20 common CNVs have been directly associated with a common disease or non-disease trait in a standard GWAS. The number of common CNVs that are candidates to explain known trait associated SNPs is not much more impressive: in an analysis of over 1500 trait-associated SNPs reported in the NHGRI GWAS database, fewer than 5% were found to be on the same genetic background as a large, common CNV.
I don’t interpret this result as evidence that large, common CNVs are not functional, but it is evidence that common CNVs are not involved in the traits that have been prioritized for genetic analysis with GWAS. Interestingly, the function of genes that tend to be included in CNV is highly non-random, and thus one may be able to create good hypotheses regarding what traits are mediated by common CNV!
Just as with rare SNPs, it seems likely that rare and unique CNVs will be more informative about disease risk.
Variation discovery without sequencing
Let’s summarize the current state of affairs. A CNV analysis with SNPs alone is likely to yield two things: (a) a set of genotypes for known, common CNVs (via imputation) that are mostly uninformative about one’s biology, and, if one has access to family data, (b) the identification of a very small number of deletions (<20) made without regard to their frequency or genomic location. If I had my parents’ data to hand, I would be excited to try (b), as it allows one to discover personal genomic variants, something otherwise impossible to do with SNP array data. It is possible to do a crude version of the parent-offspring trio approach described above just using Excel. For those who do find something interesting, be it a large and/or unreported deletion, it will give a taste of what it is like to make a scientific discovery. For those who come away empty-handed, don’t fret… that is also an authentic scientific experience!








 RSS
RSS Twitter
Twitter
There is some very limited data other than genotype (if not actual copy numbers) in the 23andme report, as detailed at SNPedia. My report contains “D”, “I”, “DD”, “II” and “DI” for deletions and insertions.
Dan, your “variation discovery without sequencing” is quite easy to do for people with parent/child data at 23andMe or Family Tree DNA, even without Excel.
At 23andMe, the Advanced Family Inheritance diagrams will show more than one segment per chromosome if the deletion is big enough. I’ve seen this happen with as few as 10,000 bases. There have been discussions about this on the 23andMe forums, usually with the term “microdeletion” rather than CNV. They do seem to occur with some regularity.
David Pike’s utility can handle data from 23andMe and Family Tree DNA. (If you are mixing data from the two companies, you may miss some microdeletions due to the limited overlap of SNPs.) He calls the parent-child discrepancies mutations, but I would attribute them to genotyping error or microdeletions.
http://www.math.mun.ca/~dapike/FF23utils/mutation.php
@ Neil — I’m not sure that Dan is referring to deletion/insertion polymorphisms (DIPs), where probes have been developed to check for those specific cases.
@Neil
Those are 23andMe’s custom probes for small indels, right? They contain things like BRCA deletions, or the NOD2 insertion. These are somewhat different from calling CNVs per se.
Right – not CNVs; thanks for clarifying their meaning.
Based on imputation from this paper
http://www.ncbi.nlm.nih.gov/pubmed/20031624
SNPedia detects one CNV today. It isn’t a particularly exciting CNV, but its a valuable test case.
Most users will find one of these
http://www.snpedia.com/index.php/Gs217
or
http://www.snpedia.com/index.php/Gs218
in their Promethease report
Perhaps also worth noting that dbVar
http://www.ncbi.nlm.nih.gov/dbvar/
the dbSNP for these sorts of variations
Very interesting post! I certainly raised an eyebrow when I first skimmed the article (before a proper read) and saw the words imputation and copy number variation being talked about so casually. Once I called myself down and actually read it start to finish I figured out what you are talking about. Very interesting premise. Thankfully the data size and complexity suggests that a simple imputation is possible in a desktop environment. We regularly do these things as part of our collaborative studies, but the data set size and complexity is on a different scale. And to impute we are usually using BEAGLE on a cluster or cloud (see http://blog.goldenhelix.com/?p=178 ).
Cheers!
Knowning more about copy number variation can be important in Prostate Cancer.
In a study in the journal Cancer Cell, the researchers analyzed the copy-number alterations in 218 cancerous prostates surgically removed at Sloan-Kettering and found that they fell into six clusters. Those clusters corresponded closely with how quickly the patients’ cancer returned.
“It was a surprise to us that so much prognostic information was there in the original samples after surgery,” Dr. Sawyers says. Ideally, “we’d be able to tell a man, ‘Your tumor looks like it’s in cluster five, so you should get surgery and radiation and perhaps even more aggressive therapy. Or, you are in cluster two, so you can relax and maybe just get another biopsy in another year and see if your cluster has changed,” he says.
It would be interesting to know what these copy number clusters were and if they could be determined by imputation from SNP data in a 23andme data download?
@ Tasker —
The full text of the article is here, but the phrase “copy-number alterations” seems to imply somatic mutations (occurring in the body after conception, and in this case, the prostate tumor itself).
http://download.cell.com/cancer-cell/pdf/PIIS1535610810002382.pdf
I am curious of your assertion:
“With a typical experiment and conservative analysis I would expect on average of 70-90 CNVs to be detected on Illumina 1M, and 20-40 CNVs with Illumina HumanHap-550”.
Having just completed the CNV discovery of the 4000 samples of the 1958 birth cohort recently (1400 of them were typed on the 550v1 and 2600 were type on the 550v3), I can say that that number of CNVs discovered per sample *on average* is only about 7. That’s quite a lot lower than your estimate.
@ Hin-Tak Leung —
Is your average number for both insertions and deletions, and is it based on signal intensity measures? If so, do you have a feel for what fraction of those would be detectable by 23andMe customers looking for discordant SNPs in parent/child pairs?
@Ann Turner:
Insertions + deletions + copy neutral LOH’s (although the last one is very low). From signal intensity measures. Probably a fair percentage (?30+ percent?) as most of them span multiple SNPs.
It is in any case a bit dubious to claim to be able to detect CNVs using one SNP – there can be so many reasons why one SNP is discordant/low/high, with or without family info, other than CNVs.
@ Hin-Tak Leung —
Sorry, I didn’t elaborate enough in my query. We’re looking at cases where there are several discordant SNPs within a few thousand kb of each other, embedded in a run of homozygous SNPs. Would you say that’s a reliable method?
@Ann Turner
A few thousand *kb* (i.e. a few Mb)?
discordant SNPs in parent/child pairs can be genotyping errors – it is only when they happens together (in neighbouring SNPs) where one may say they aren’t. OTOH, what’s the purpose of such detection? Many ins/dels have no health consequences.
@ Hin-Tak Leung —
No, I did mean kb. A concrete example in my files: 11 SNPs covering a span of 110,000 bases, and six of are discordant. To be sure, larger deletions are bound to have more consequences, but we’re exploring ways to look at our own data.
I have a three year old boy that has been diagnosed with language disorders, lack of coordination, eustachian tube dysfunction with no ear infection history and his comprehension is very weak for a child of his age. My son had an EEG done with normal results,an MRI with normal results, fragile x results were also normal he also had an Micro-array study and the interpretation was Copy Number Variation Identified: arr 5p15.2(12,669,534-12,785,251)x1
Should we the parents get genetic testing? Are there any more testing for my child, so we can figure out what is the problem? we would like to give him the help he needs or should I just relax because this result is not something for me to worry. His neurologist said that he is going to be a little different in comparison to other children and I should research on the net for new articles about copy number variants and maybe in five years retest him.