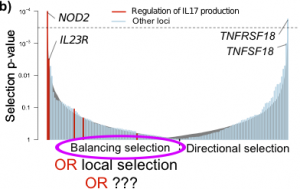
 As I mentioned a few weeks ago, we recently published a large study into the genetics of inflammatory bowel disease (IBD), which included a number of analyses digging into the biology and evolutionary history of IBD genetic risk. Gratifyingly, our paper has stimulated a lot of discussion among other scientists, which has generated several ideas about future directions for this work. One question that was raised by several population-genetics experts at ASHG was about our natural selection analysis, and in particular our claim to discover an enrichment of balancing selection in IBD loci. In the paper, we found clear signals of natural selection on IBD loci, a subset of which we interpreted as balancing selection. In this post I will set out how I came to this conclusion, but then outline another explanation that could explain the results: recent local positive selection in Europeans.
As I mentioned a few weeks ago, we recently published a large study into the genetics of inflammatory bowel disease (IBD), which included a number of analyses digging into the biology and evolutionary history of IBD genetic risk. Gratifyingly, our paper has stimulated a lot of discussion among other scientists, which has generated several ideas about future directions for this work. One question that was raised by several population-genetics experts at ASHG was about our natural selection analysis, and in particular our claim to discover an enrichment of balancing selection in IBD loci. In the paper, we found clear signals of natural selection on IBD loci, a subset of which we interpreted as balancing selection. In this post I will set out how I came to this conclusion, but then outline another explanation that could explain the results: recent local positive selection in Europeans.
Continue reading ‘Looking closer at natural selection in inflammatory bowel disease’
 Out in Nature this week is a paper by three Genomes Unzipped authors reporting 71 new genetic associations with inflammatory bowel disease (IBD). This breaks the record for the largest number of associations for any common disease, and includes many new and interesting biological insights that you should all go and read about in the paper itself (pay-to-access I’m afraid) or on the Sanger Institute’s website.
Out in Nature this week is a paper by three Genomes Unzipped authors reporting 71 new genetic associations with inflammatory bowel disease (IBD). This breaks the record for the largest number of associations for any common disease, and includes many new and interesting biological insights that you should all go and read about in the paper itself (pay-to-access I’m afraid) or on the Sanger Institute’s website. A paper out in PLoS Genetics this week takes a step towards using genome-wide association data to reconstruct functional pathways. Using protein-protein interaction data and tissue-specific expression data, the authors reconstruct biochemical pathways that underlie various diseases, by looking for variants that interact with genes in GWAS regions. These networks can then tell us about what systems are disrupted by GWAS variants as a whole, as well as identifying potential drug targets. The figure to the right shows the network constructed for Crohn’s disease; large colored circles are genes in GWAS loci, small grey circles are other genes in the network they constructed. As an interesting side note, the GWAS variants were taken from a 2008 study; since then, we have published a new meta-analysis, which implicated a lot of new regions. 10 genes in these regions, marked as small red circles on the figure, were also in the disease network. [LJ]
A paper out in PLoS Genetics this week takes a step towards using genome-wide association data to reconstruct functional pathways. Using protein-protein interaction data and tissue-specific expression data, the authors reconstruct biochemical pathways that underlie various diseases, by looking for variants that interact with genes in GWAS regions. These networks can then tell us about what systems are disrupted by GWAS variants as a whole, as well as identifying potential drug targets. The figure to the right shows the network constructed for Crohn’s disease; large colored circles are genes in GWAS loci, small grey circles are other genes in the network they constructed. As an interesting side note, the GWAS variants were taken from a 2008 study; since then, we have published a new meta-analysis, which implicated a lot of new regions. 10 genes in these regions, marked as small red circles on the figure, were also in the disease network. [LJ] There is a real “wow” paper out in pre-print at the journal Genetics in Medicine. It is a wonderful example of the application of cutting edge sequencing technology to solve a medical mystery. Even better, the authors also include an auxiliary discussion about the medical and ethical issues surrounding the diagnosis, which raises some interesting issues about the transition from research to clinical sequencing.
There is a real “wow” paper out in pre-print at the journal Genetics in Medicine. It is a wonderful example of the application of cutting edge sequencing technology to solve a medical mystery. Even better, the authors also include an auxiliary discussion about the medical and ethical issues surrounding the diagnosis, which raises some interesting issues about the transition from research to clinical sequencing. RSS
RSS Twitter
Twitter
Recent Comments